3.2. Example 1 - Crowded with offset to sky - Using the “reduce” command line
This chapter will guide you on reducing GSAOI data using command line tools. In this example we reduce a GSAOI observation of the resolved outskirt of a nearby galaxy. The observation is a dither-on-target with offset-to-sky sequence. Just open a terminal to get started.
While the example cannot possibly cover all situations, it will help you get acquainted with the reduction of GSAOI data with DRAGONS. We encourage you to look at the Tips and Tricks and Issues and Limitations chapters to learn more about GSAOI data reduction.
DRAGONS installation comes with a set of scripts that are used to reduce astronomical data. The most important script is called “reduce”, which is extensively explained in the Recipe System Users Manual. It is through that command that a DRAGONS reduction is launched.
For this tutorial, we will be also using the following supplemental tools: “dataselect”, “showd”, “typewalk”, and “caldb”.
3.2.1. The dataset
If you have not already, download and unpack the tutorial’s data package. Refer to Downloading the tutorial datasets for the links and simple instructions.
The dataset specific to this example is described in:
Here is a copy of the table for quick reference.
Science |
S20170505S0095-110
|
Kshort-band, on target, 60 s
|
Flats |
S20170505S0030-044
S20170505S0060-074
|
Lamp on, Kshort, for science
Lamp off, Kshort, for science
|
Standard star |
S20170504S0114-117
|
Kshort, standard star, 30 s
|
BMP |
bpm_20121104_gsaoi_gsaoi_11_full_4amp.fits
|
|
Note
A master dark is not needed for GSAOI. The dark current is very low.
3.2.2. Set up the Calibration Service
Important
Remember to set up the calibration service.
Instructions to configure and use the calibration service are found in Setting up the Calibration Service, specifically the these sections: The Configuration File and Usage from the Command Line.
3.2.3. Check files
For this example, all the raw files we need are in the same directory called
../playdata/example1. Let us learn a bit about the data we have.
Ensure that you are in the playground directory and that the conda
environment that includes DRAGONS has been activated.
Let us call the command tool “typewalk”:
$ typewalk -d ../playdata/example1
directory: /data/workspace/gsaoiimg_tutorial/playdata/example1
S20170504S0114.fits ............... (GEMINI) (GSAOI) (IMAGE) (RAW) (SIDEREAL) (SOUTH) (UNPREPARED)
...
S20170505S0030.fits ............... (AZEL_TARGET) (CAL) (DOMEFLAT) (FLAT) (GEMINI) (GSAOI) (IMAGE) (LAMPON) (NON_SIDEREAL) (RAW) (SOUTH) (UNPREPARED)
...
S20170505S0060.fits ............... (AZEL_TARGET) (CAL) (DOMEFLAT) (FLAT) (GEMINI) (GSAOI) (IMAGE) (LAMPOFF) (NON_SIDEREAL) (RAW) (SOUTH) (UNPREPARED)
...
S20170505S0095.fits ............... (GEMINI) (GSAOI) (IMAGE) (RAW) (SIDEREAL) (SOUTH) (UNPREPARED)
...
S20170505S0110.fits ............... (GEMINI) (GSAOI) (IMAGE) (RAW) (SIDEREAL) (SOUTH) (UNPREPARED)
Done DataSpider.typewalk(..)
This command will open every FITS file within the folder passed after the -d
flag (recursively) and will print an unsorted table with the file names and the
associated tags. For example, calibration files will always have the CAL
tag. Flat images will always have the FLAT tag. This means that we can
start getting to know a bit more about our data set just by looking at the tags.
The output above was trimmed for presentation.
3.2.4. Create File lists
This data set contains science and calibration frames. For some program, it could have different observed targets and different exposure times depending on how you like to organize your raw data.
The DRAGONS data reduction pipeline does not organize the data for you. You have to do it. DRAGONS provides tools to help you with that.
The first step is to create input file lists. The tool “dataselect” helps with that. It uses Astrodata tags and “descriptors” to select the files and send the filenames to a text file that can then be fed to “reduce”. (See the Astrodata User Manual for information about Astrodata.)
First, navigate to the playground directory in the unpacked data package:
cd <path>/gsaoiim_tutorial/playground
3.2.4.1. A list for the flats
Let us create the list containing the domeflats:
$ dataselect --tags FLAT ../playdata/example1/*.fits -o flats_Kshort.list
We know that our dataset has only one filter (Kshort). If our dataset
contained data with more filters, we would have had to use the --expr
option to select the appropriate filter as follows:
$ dataselect --tags FLAT --expr "filter_name=='Kshort'" ../playdata/example1/*.fits -o flats_Kshort.list
Note
To see the name of the filter, use “showd” (show descriptor):
$ showd ../playdata/example1/*.fits -d filter_name
-------------------------------------------------------------
filename filter_name
-------------------------------------------------------------
../playdata/example1/S20170504S0114.fits Kshort_G1105&Clear
...
...
3.2.4.2. A list for the standard star
In this case we have only one standard star. Indeed, we can confirm that by selecting on partner calibrations and showing the object name:
$ dataselect --expr 'observation_class=="partnerCal"' ../playdata/example1/*.fits | showd -d object
-------------------------------------------------
filename object
-------------------------------------------------
../playdata/example1/S20170504S0114.fits 9132
../playdata/example1/S20170504S0115.fits 9132
../playdata/example1/S20170504S0116.fits 9132
../playdata/example1/S20170504S0117.fits 9132
If we had more than one object, a list for each standard star is created by
using the object descriptor as a selection criterium in “dataselect”:
$ dataselect --expr 'object=="9132"' ../playdata/example1/*.fits -o std_9132.list
3.2.4.3. A list for the science observations
The rest is the data with your science target. Before we create a new list, let us check that indeed we have only one science target and a unique exposure time:
$ dataselect --expr 'observation_class=="science"' ../playdata/example1/*.fits | showd -d object,exposure_time
------------------------------------------------------------------
filename object exposure_time
------------------------------------------------------------------
../playdata/example1/S20170505S0095.fits NGC5128 60.0
../playdata/example1/S20170505S0096.fits NGC5128 60.0
...
../playdata/example1/S20170505S0109.fits NGC5128 60.0
../playdata/example1/S20170505S0110.fits NGC5128 60.0
Just to demonstrate how expression are built, let us consider that we need to
select only the files for which object is NGC5128 and exposure_time
is 60 seconds. We also want to pass the output to a new list:
$ dataselect --expr '(observation_class=="science" and exposure_time==60.)' ../playdata/example1/*.fits -o science.list
3.2.5. Bad Pixel Mask
Starting with DRAGONS v3.1, the bad pixel masks (BPMs) are now handled as calibrations. They are downloadable from the archive instead of being packaged with the software. They are automatically associated like any other calibrations. This means that the user now must download the BPMs along with the other calibrations and add the BPMs to the local calibration manager.
See Getting Bad Pixel Masks from the archive in Tips and Tricks to learn about the various ways to get the BPMs from the archive.
To add the static BPM included in the data package to the local calibration database:
caldb add ../playdata/example1/bpm*.fits
3.2.6. Create a Master Flat Field
The GSAOI Kshort master flat is created from a series of lamp-on and lamp-off dome exposures. They should all have the same exposure time. Each flavor is stacked (averaged), then the lamp-off stack is subtracted from the lamp-on stack and the result normalized.
We create the master flat field and add it to the calibration manager as follows:
$ reduce @flats_Kshort.list
The @ character before the name of the input file is the “at-file” syntax.
More details can be found in the "at-file" Facility documentation.
Because the database was given the “store” option in the dragonsrc file,
the processed dark will be automatically added to the database at the end of
the recipe.
Note
The file name of the output processed flat is the file name of the
first file in the list with _flat appended as a suffix. This the
general naming scheme used by “reduce”.
Note
- If you wish to inspect the processed calibrations before adding them
to the calibration database, remove the “store” option attached to the database in the
dragonsrcconfiguration file. You will then have to add the calibrations manually following your inspection, eg.
caldb add S20170505S0030_flat.fits
Note
The master flat will be saved in the same folder where reduce was
called and inside the ./calibrations/processed_flat folder. The latter
location is to cache a copy of the file. This applies to all the processed
calibration.
Here is an example of a master flat:
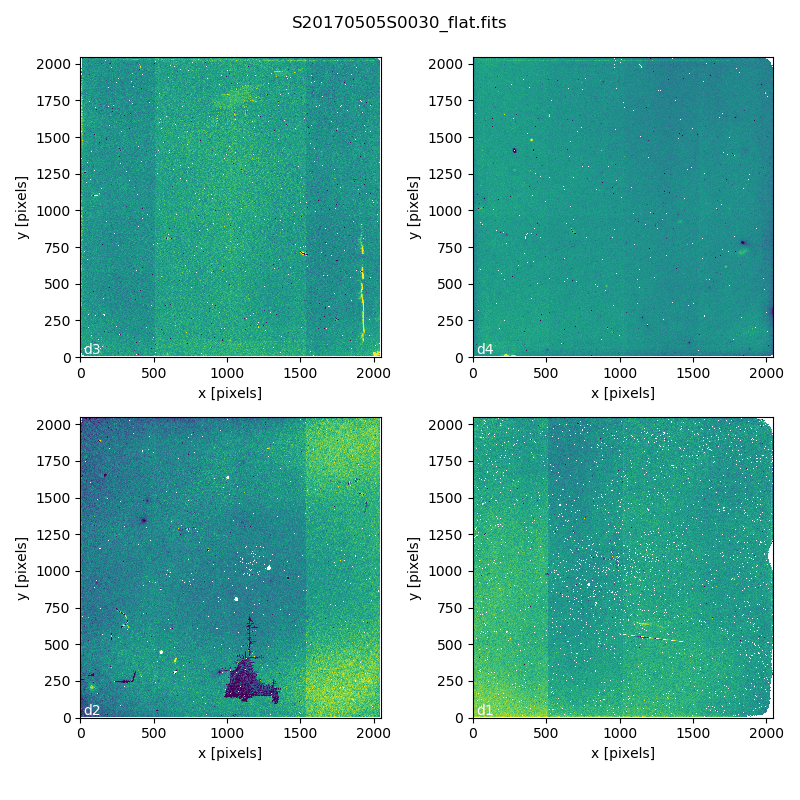
Master Flat - K-Short Band
Note that this figure shows the masked pixels in white color but not all the detector features are masked. For example, the “Christmas Tree” on detector 2 can be easily noticed but was not masked.
3.2.7. Reduce Standard Star
The standard star is reduced essentially the same way as the science target (next section). The processed flat field that we added earlier to the local calibration database will be fetched automatically. Also, in this case the standard star was obtained using ROIs (Regions-of-Interest) which do not match the flat field. The software will recognize that the flat field is still valid and will crop it to match the ROIs.
$ reduce @std_9132.list
Note
The reduce command will automatically align and stack the images.
Therefore, it is no longer necessary to use the disco_stu tool for
GSAOI data.
3.2.8. Reduce the Science Images
This is an observation of a galaxy with offset to sky. We need to turn off the additive offsetting of the sky because the target fills the field of view and does not represent a reasonable sky background. If the offsetting is not turned off in this particular case, it results in an over-subtraction of the sky frame.
Note
Unlike the other near-IR instruments, the additive offset_sky
parameter is used by default to adjust the sky frame background for
GSAOI instead of the multiplicative scale_sky parameter. It was
found to work better when the sky background per pixel is very low,
which is common due to the short exposure time needed to avoid
saturating stars and the small pixel scale. The reader is encouraged
to experiment with scale_sky if offset_sky does not seem to
lead to an optimal sky subtraction.
(Remember that when the source is extended, both parameters normally need to be turned off.)
The sky frame comes from off-target sky observations. We feed the pipeline all the on-target and off-target frames. The software will split the on-target and the off-target appropriately using information in the headers.
Once we have our calibration files processed and added to the database, ready
for retrieval, we can run reduce on our science data.
$ reduce @science.list -p skyCorrect:offset_sky=False
This command will generate flat corrected files, align them,
stack them, and orient them such that North is up and East is left. The final
image will have the name of the first file in the set, with the suffix
_image. The on-target files are the ones that have been flat corrected
(_flatCorrected), and scaled (_countsScaled). There should be nine
of these.
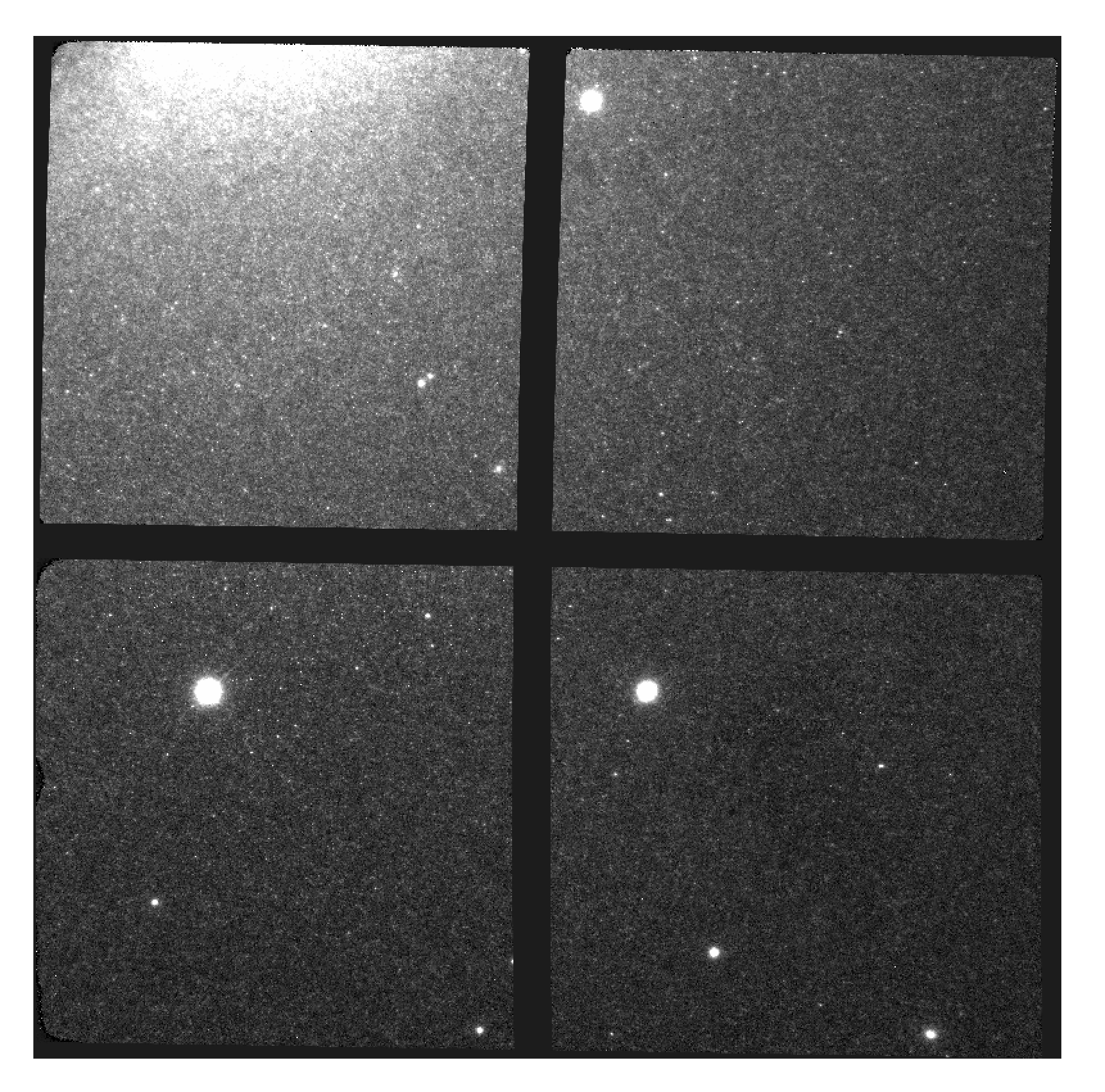

S20170505S0095 - Final flat corrected, aligned, and stacked image
The figure above shows the final flat-corrected, aligned, and stacked frame.
For absolute distortion correction and astrometry, reduce can use a
reference catalog provided by the user. Without a reference catalog, like
above, only the relative distortion between the frames is accounted for.
The output stack units are in electrons (header keyword BUNIT=electrons). The output stack is stored in a multi-extension FITS (MEF) file. The science signal is in the “SCI” extension, the variance is in the “VAR” extension, and the data quality plane (mask) is in the “DQ” extension.